Note on performance
In section 3-1, the use of the SOAP descriptor in GPR can be computationally intensive.
On high-performance computing (HPC) or GPU systems, the SOAP-based GPR runs significantly faster.
In this Colab environment, you can switch to a GPU by going to
Runtime → Change runtime type → Hardware accelerator → T4 GPU
try:
import torch
print("successfully imported torch")
print(torch.__version__)
except ImportError:
!! pip install torch torchvision torchaudio --user --index-url https://download.pytorch.org/whl/cpu
print("completed installing torch")successfully imported torch
2.6.0+cu124
try:
import numpy as np
print("successfully imported numpy")
print(np.__version__)
except ImportError:
!! pip install numpy --user --upgrade
print("completed installing numpy")successfully imported numpy
1.26.4
try:
from matplotlib import pyplot as plt
print("successfully imported matplotlib")
except ImportError:
!! pip install matplotlib --user --upgrade
print("completed installing matplotlib")successfully imported matplotlib
!! pip install aenet-gpr --user --upgrade
print("completed installing aenet-gpr")
! pip show aenet-gprcompleted installing aenet-gpr
Name: aenet-gpr
Version: 2.6.5
Summary: Atomistic simulation tools based on Gaussian Processes Regression
Home-page: https://github.com/atomisticnet/aenet-gpr
Author: In Won Yeu
Author-email: iy2185@columbia.edu
License: MPL-2.0
Location: /data/home/iy2185/.local/lib/python3.12/site-packages
Requires: ase
Required-by:
import os, sys, site
sys.path.append(site.USER_SITE)
import aenet_gpr
print(aenet_gpr.__version__)2.6.5
from IPython.display import Image
try:
import ase.io
print("successfully imported ase")
except ImportError:
!! pip install ase --user --upgrade
print("completed installing ASE")
import ase.io
print("successfully imported ase")successfully imported ase
1. Prepare training/test data¶
We are using train/test HSE06 data of Li/EC interface prepared in aenet-gpr/example/3_Li-EC/
import os.path
! rm -rf 3_Li-EC
! mkdir 3_Li-EC
if os.path.isfile("../example/3_Li-EC/train_set_tutorial.zip"):
! unzip -oq ../example/3_Li-EC/train_set_tutorial.zip -d ./3_Li-EC/
print("number of train data:")
! find ./3_Li-EC/train_set_tutorial/ -type f | wc -l
else:
! wget https://github.com/atomisticnet/aenet-gpr/raw/refs/heads/main/example/3_Li-EC/train_set_tutorial.zip
! unzip -oq train_set_tutorial.zip -d ./3_Li-EC/
! rm train_set_tutorial.zip
print("number of train data:")
! find ./3_Li-EC/train_set_tutorial/ -type f | wc -l--2025-10-19 17:05:28-- https://github.com/atomisticnet/aenet-gpr/raw/refs/heads/main/example/3_Li-EC/train_set_tutorial.zip
Resolving github.com (github.com)... 140.82.112.3
Connecting to github.com (github.com)|140.82.112.3|:443... connected.
HTTP request sent, awaiting response... 302 Found
Location: https://raw.githubusercontent.com/atomisticnet/aenet-gpr/refs/heads/main/example/3_Li-EC/train_set_tutorial.zip [following]
--2025-10-19 17:05:28-- https://raw.githubusercontent.com/atomisticnet/aenet-gpr/refs/heads/main/example/3_Li-EC/train_set_tutorial.zip
Resolving raw.githubusercontent.com (raw.githubusercontent.com)... 185.199.110.133, 185.199.109.133, 185.199.108.133, ...
Connecting to raw.githubusercontent.com (raw.githubusercontent.com)|185.199.110.133|:443... connected.
HTTP request sent, awaiting response... 200 OK
Length: 74855 (73K) [application/zip]
Saving to: ‘train_set_tutorial.zip’
train_set_tutorial. 100%[===================>] 73.10K --.-KB/s in 0.001s
2025-10-19 17:05:28 (52.6 MB/s) - ‘train_set_tutorial.zip’ saved [74855/74855]
number of train data:
30
if os.path.isfile("../example/3_Li-EC/test_set_tutorial.zip"):
! unzip -oq ../example/3_Li-EC/test_set_tutorial.zip -d ./3_Li-EC/
print("number of test data:")
! find ./3_Li-EC/test_set_tutorial/ -type f | wc -l
else:
! wget https://github.com/atomisticnet/aenet-gpr/raw/refs/heads/main/example/3_Li-EC/test_set_tutorial.zip
! unzip -oq test_set_tutorial.zip -d ./3_Li-EC/
! rm test_set_tutorial.zip
print("number of test data:")
! find ./3_Li-EC/test_set_tutorial/ -type f | wc -l--2025-10-19 17:05:28-- https://github.com/atomisticnet/aenet-gpr/raw/refs/heads/main/example/3_Li-EC/test_set_tutorial.zip
Resolving github.com (github.com)... 140.82.112.3
Connecting to github.com (github.com)|140.82.112.3|:443... connected.
HTTP request sent, awaiting response... 302 Found
Location: https://raw.githubusercontent.com/atomisticnet/aenet-gpr/refs/heads/main/example/3_Li-EC/test_set_tutorial.zip [following]
--2025-10-19 17:05:29-- https://raw.githubusercontent.com/atomisticnet/aenet-gpr/refs/heads/main/example/3_Li-EC/test_set_tutorial.zip
Resolving raw.githubusercontent.com (raw.githubusercontent.com)... 185.199.110.133, 185.199.109.133, 185.199.108.133, ...
Connecting to raw.githubusercontent.com (raw.githubusercontent.com)|185.199.110.133|:443... connected.
HTTP request sent, awaiting response... 200 OK
Length: 149388 (146K) [application/zip]
Saving to: ‘test_set_tutorial.zip’
test_set_tutorial.z 100%[===================>] 145.89K --.-KB/s in 0.003s
2025-10-19 17:05:29 (41.8 MB/s) - ‘test_set_tutorial.zip’ saved [149388/149388]
number of test data:
60
atoms = ase.io.read("./3_Li-EC/test_set_tutorial/file_0000.xsf")
ase.io.write('Li-EC.png', atoms)
Image("Li-EC.png")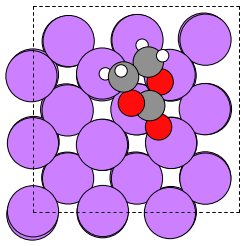
2-1. Train–Test with default Cartesian fingerprint¶
! rm -f train.in
if os.path.isfile("../example/3_Li-EC/train.in"):
! cp ../example/3_Li-EC/train.in .
else:
! wget https://github.com/atomisticnet/aenet-gpr/raw/refs/heads/main/example/3_Li-EC/train.in--2025-10-19 17:05:30-- https://github.com/atomisticnet/aenet-gpr/raw/refs/heads/main/example/3_Li-EC/train.in
Resolving github.com (github.com)... 140.82.112.3
Connecting to github.com (github.com)|140.82.112.3|:443... connected.
HTTP request sent, awaiting response... 302 Found
Location: https://raw.githubusercontent.com/atomisticnet/aenet-gpr/refs/heads/main/example/3_Li-EC/train.in [following]
--2025-10-19 17:05:30-- https://raw.githubusercontent.com/atomisticnet/aenet-gpr/refs/heads/main/example/3_Li-EC/train.in
Resolving raw.githubusercontent.com (raw.githubusercontent.com)... 185.199.110.133, 185.199.109.133, 185.199.108.133, ...
Connecting to raw.githubusercontent.com (raw.githubusercontent.com)|185.199.110.133|:443... connected.
HTTP request sent, awaiting response... 200 OK
Length: 587 [text/plain]
Saving to: ‘train.in’
train.in 100%[===================>] 587 --.-KB/s in 0s
2025-10-19 17:05:30 (10.8 MB/s) - ‘train.in’ saved [587/587]
! cat train.in# File path
Train_file ./3_Li-EC/train_set_tutorial/file_*.xsf
Test_file ./3_Li-EC/test_set_tutorial/file_*.xsf
# Train model save (default: False)
# Train_model_save True
# File format (default: xsf)
File_format xsf
# Descriptor (default: cartesian coordinates)
Descriptor cart
# Kernel parameter
# scale 0.4
# weight 1.0
# Data process (default: batch, 25)
data_process batch
batch_size 25
# Flags for xsf file writing (default: False)
Train_write False
Test_write False
# Additional_write True
# Data augmentation parameter (default: 0.055, 25)
# Disp_length 0.05
# Num_copy 3
! python -m aenet_gpr train.in======================================================================
aenet-GPR: surrogate GPR for GPR-ANN indirect force training
======================================================================
2025-10-19 17:05:34.
Developed by In Won Yeu
This program performs three main steps:
1. Train: Generates a GPR model using the provided structure, energy, and force data.
2. Test: Uses the generated GPR model to predict values for the test set structures.
3. Augmentation: Performs data augmentation in xsf file format, compatible with aenet-(PyTorch),
supporting a GPR-ANN training in conjunction with aenet-(PyTorch).
Each of these steps is executed once the input file (train.in) contains the keywords:
Train_file [train file path]
Test_file [test file path]
Additional_write [True]
======================================================================
Train
======================================================================
Read reference training data
Time needed for reading data: 0.077833 s
Maximum CPU memory used: 0.411930 GB
Maximum GPU memory used: 0.000000 GB
Energy data size: (30,) # (N_data, )
Force data size: (30, 46, 3) # (N_data, N_atom, 3)
----------------------------------------------------------------------
Candidate scales: tensor([0.2000, 0.4000, 0.8000])
Updated scale: 0.800000011920929
Updated weight: 1.1156722220085324
----------------------------------------------------------------------
Model train
Training parameters
{'num_data': 30, 'calculator': {'kerneltype': 'sqexp', 'scale': tensor(0.8000), 'weight': tensor(1.1157, dtype=torch.float64), 'noise': tensor(1.0000e-06, dtype=torch.float64), 'noisefactor': tensor(0.5000, dtype=torch.float64), 'prior': tensor(-16685.0192, dtype=torch.float64)}, 'fix_ind': None, 'pbc': True, 'species': ['Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'C', 'C', 'O', 'H', 'H', 'O', 'H', 'H', 'C', 'O'], 'num_atom': 46}
Time needed for training: 3.383716 s
Maximum CPU memory used: 0.997082 GB
Maximum GPU memory used: 0.000000 GB
----------------------------------------------------------------------
======================================================================
Test
======================================================================
----------------------------------------------------------------------
Model evaluation for test set
Time needed for test evaluation: 4.836310 s
Maximum CPU memory used: 1.618618 GB
Maximum GPU memory used: 0.000000 GB
----------------------------------------------------------------------
GPR energy MAE (eV): 0.1636123145555151
GPR force MAE (eV/Ang): 0.5956040013558996
Saving test target to [energy_test_reference.npy] and [force_test_reference.npy]
Saving GPR prediction to [energy_test_gpr.npy], [force_test_gpr.npy], [unc_e_test_gpr.npy], and [unc_f_test_gpr.npy]
2-2. Visualize the results¶
energy_test_gpr = np.load("./energy_test_gpr.npy")
energy_test_reference = np.load("./energy_test_reference.npy")
force_test_gpr = np.load("./force_test_gpr.npy")
force_test_reference = np.load("./force_test_reference.npy")
unc_e_test_gpr = np.load("./unc_e_test_gpr.npy")
unc_f_test_gpr = np.load("./unc_f_test_gpr.npy")diff_per_atom = (energy_test_reference - energy_test_gpr) / 46
print(diff_per_atom.min(), diff_per_atom.max())-0.007817935342324705 0.012918689117372118
fig = plt.figure(figsize=(7, 6))
ax2 = fig.subplots()
font_x = {'size': 20, 'color': 'black'}
font_y = {'size': 20, 'color': 'black'}
font_tick = {'size': 16, 'color': 'black'}
# The second plot
counts, bins = np.histogram(diff_per_atom, bins=30, range=(-0.005, 0.05))
ax2.hist(bins[:-1], bins, weights=counts, facecolor='dimgray', alpha=1.0, edgecolor='black')
ax2.set_xlabel("DFT $-$ GPR (eV/atom)", fontdict=font_x)
ax2.set_xlim([-0.03, 0.03])
x_labels = [round(label, 2) for label in ax2.get_xticks()]
ax2.set_xticks(x_labels)
ax2.set_xlim([-0.03, 0.03])
ax2.set_xticklabels(x_labels, fontdict=font_tick)
ax2.set_ylabel("Number of structures", fontdict=font_y)
ax2.set_ylim([0, 25])
y_labels = [int(label) for label in ax2.get_yticks()]
ax2.set_yticks(y_labels)
ax2.set_ylim([0, 25])
ax2.set_yticklabels(y_labels, fontdict=font_tick)
[x.set_linewidth(1.5) for x in ax2.spines.values()]
ax2.tick_params(bottom=False, top=True, left=True, right=True)
ax2.tick_params(labelbottom=True, labeltop=False, labelleft=True, labelright=False)
ax2.tick_params(direction='in', length=8, width=1.5)
ax2.grid(True, which='both', linestyle=':', linewidth=0.7, alpha=0.7)
fig.tight_layout()
plt.show()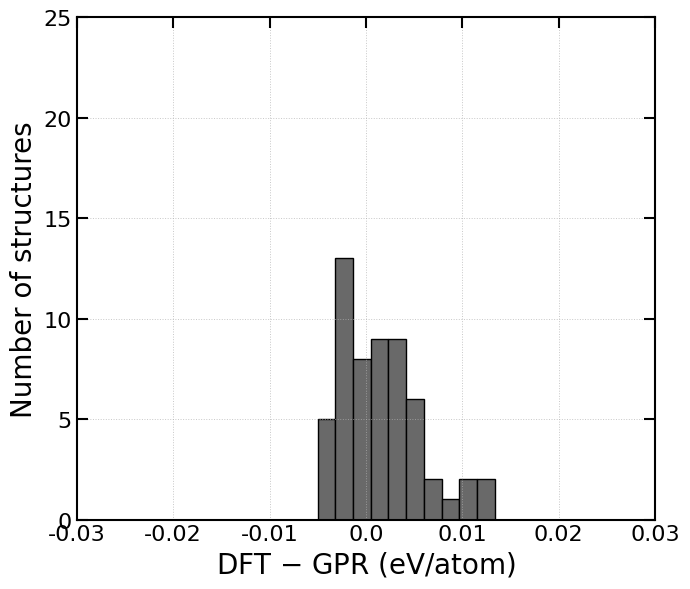
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import pearsonr
# --- Force error vs uncertainty ---
force_error = np.abs(force_test_gpr - force_test_reference).reshape(-1)
unc_f = unc_f_test_gpr.reshape(-1)
r_force, _ = pearsonr(force_error, unc_f)
plt.figure(figsize=(6, 5))
plt.scatter(unc_f, force_error, alpha=0.6, edgecolors='k')
plt.xlabel("Force uncertainty (eV/Å)")
plt.ylabel("Force absolute error (eV/Å)")
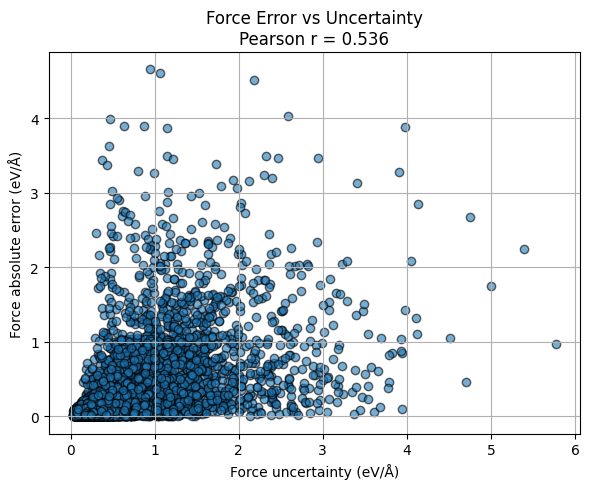
plt.title(f"Force Error vs Uncertainty\nPearson r = {r_force:.3f}")
plt.grid(True)
# plt.xlim(0.0, 1.0)
# plt.ylim(0, 2.0)
plt.tight_layout()
plt.show()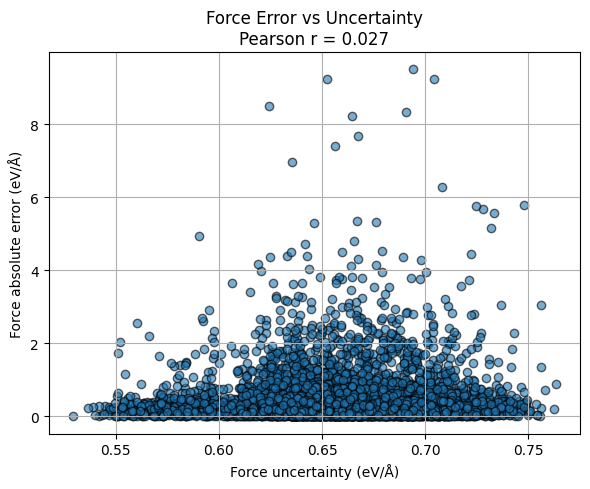
3-1. Train–Test–Augment with SOAP fingerprint¶
! rm -f train_soap.in
if os.path.isfile("../example/3_Li-EC/train_soap.in"):
! cp ../example/3_Li-EC/train_soap.in .
else:
! wget https://github.com/atomisticnet/aenet-gpr/raw/refs/heads/main/example/3_Li-EC/train_soap.in--2025-10-19 17:07:20-- https://github.com/atomisticnet/aenet-gpr/raw/refs/heads/main/example/3_Li-EC/train_soap.in
Resolving github.com (github.com)... 140.82.112.3
Connecting to github.com (github.com)|140.82.112.3|:443... connected.
HTTP request sent, awaiting response... 302 Found
Location: https://raw.githubusercontent.com/atomisticnet/aenet-gpr/refs/heads/main/example/3_Li-EC/train_soap.in [following]
--2025-10-19 17:07:20-- https://raw.githubusercontent.com/atomisticnet/aenet-gpr/refs/heads/main/example/3_Li-EC/train_soap.in
Resolving raw.githubusercontent.com (raw.githubusercontent.com)... 185.199.110.133, 185.199.109.133, 185.199.108.133, ...
Connecting to raw.githubusercontent.com (raw.githubusercontent.com)|185.199.110.133|:443... connected.
HTTP request sent, awaiting response... 200 OK
Length: 1077 (1.1K) [text/plain]
Saving to: ‘train_soap.in’
train_soap.in 100%[===================>] 1.05K --.-KB/s in 0s
2025-10-19 17:07:20 (18.1 MB/s) - ‘train_soap.in’ saved [1077/1077]
! cat train_soap.in# File path
Train_file ./3_Li-EC/train_set_tutorial/file_*.xsf
Test_file ./3_Li-EC/test_set_tutorial/file_*.xsf
# File format (default: xsf)
File_format xsf
# Descriptor (default: cartesian coordinates)
Descriptor soap
# Soap parameters
soap_r_cut 5.0
soap_n_max 6
soap_l_max 4
soap_centers [[2.20113706670393, 2.328998192856251, 6.952547732109352], [2.20113706670393, 2.328998192856251, 11.895790642109352], [2.20113706670393, 6.760484232856251, 6.952547732109352], [2.20113706670393, 6.760484232856251, 11.895790642109352], [6.63924050670393, 2.328998192856251, 6.952547732109352], [6.63924050670393, 2.328998192856251, 11.895790642109352], [6.63924050670393, 6.760484232856251, 6.952547732109352], [6.63924050670393, 6.760484232856251, 11.895790642109352]]
soap_n_jobs 32
# Kernel parameter
# scale 0.4
# weight 1.0
# Data process (default: batch, 25)
data_process batch
batch_size 10
# Flags for xsf file writing (default: False)
Train_write False
Test_write False
Additional_write True
# Data augmentation parameter (default: 0.055, 25)
Disp_length 0.05
Num_copy 2
! rm -rf additional_xsf/
! python -m aenet_gpr train_soap.in======================================================================
aenet-GPR: surrogate GPR for GPR-ANN indirect force training
======================================================================
2025-10-19 17:07:24.
Developed by In Won Yeu
This program performs three main steps:
1. Train: Generates a GPR model using the provided structure, energy, and force data.
2. Test: Uses the generated GPR model to predict values for the test set structures.
3. Augmentation: Performs data augmentation in xsf file format, compatible with aenet-(PyTorch),
supporting a GPR-ANN training in conjunction with aenet-(PyTorch).
Each of these steps is executed once the input file (train.in) contains the keywords:
Train_file [train file path]
Test_file [test file path]
Additional_write [True]
======================================================================
Train
======================================================================
Read reference training data
Time needed for reading data: 0.076803 s
Maximum CPU memory used: 0.411224 GB
Maximum GPU memory used: 0.000000 GB
Energy data size: (30,) # (N_data, )
Force data size: (30, 46, 3) # (N_data, N_atom, 3)
----------------------------------------------------------------------
Candidate scales: tensor([0.2000, 0.4000, 0.8000])
Updated scale: 0.800000011920929
Updated weight: 0.7879021148110678
----------------------------------------------------------------------
Model train
Training parameters
{'num_data': 30, 'calculator': {'kerneltype': 'sqexp', 'scale': tensor(0.8000), 'weight': tensor(0.7879, dtype=torch.float64), 'noise': tensor(1.0000e-06, dtype=torch.float64), 'noisefactor': tensor(0.5000, dtype=torch.float64), 'prior': tensor(-16684.1882, dtype=torch.float64)}, 'fix_ind': None, 'pbc': True, 'species': ['Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'C', 'C', 'O', 'H', 'H', 'O', 'H', 'H', 'C', 'O'], 'num_atom': 46}
Time needed for training: 135.652588 s
Maximum CPU memory used: 3.810555 GB
Maximum GPU memory used: 0.000000 GB
----------------------------------------------------------------------
======================================================================
Test
======================================================================
----------------------------------------------------------------------
Model evaluation for test set
Time needed for test evaluation: 89.152006 s
Maximum CPU memory used: 3.903370 GB
Maximum GPU memory used: 0.000000 GB
----------------------------------------------------------------------
GPR energy MAE (eV): 0.1501698888436901
GPR force MAE (eV/Ang): 0.3222749835025399
Saving test target to [energy_test_reference.npy] and [force_test_reference.npy]
Saving GPR prediction to [energy_test_gpr.npy], [force_test_gpr.npy], [unc_e_test_gpr.npy], and [unc_f_test_gpr.npy]
======================================================================
Augmentation
======================================================================
----------------------------------------------------------------------
Additional structure generation
Time needed for additional generation: 0.012513 s
Maximum CPU memory used: 3.903370 GB
Maximum GPU memory used: 0.000000 GB
Displacement length (Ang): 0.05
Multiple: 2
N_additional = N_reference (30) * Multiple (2) = 60
----------------------------------------------------------------------
----------------------------------------------------------------------
Model evaluation for additional set
Test parameters
{'num_data': 60, 'calculator': {'kerneltype': 'sqexp', 'scale': tensor(0.8000), 'weight': tensor(0.7879, dtype=torch.float64), 'noise': tensor(1.0000e-06, dtype=torch.float64), 'noisefactor': tensor(0.5000, dtype=torch.float64), 'prior': tensor(-16684.1882, dtype=torch.float64)}, 'fix_ind': None, 'pbc': True, 'species': ['Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'Li', 'C', 'C', 'O', 'H', 'H', 'O', 'H', 'H', 'C', 'O'], 'num_atom': 46}
Time needed for additional evaluation: 88.011260 s
Maximum CPU memory used: 3.940384 GB
Maximum GPU memory used: 0.000000 GB
----------------------------------------------------------------------
Saving GPR prediction to [energy_additional_gpr.npy], [force_additional_gpr.npy], [unc_e_additional_gpr.npy], and [unc_f_additional_gpr.npy]
======================================================================
Writing additional xsf files to additional_xsf
Time needed for writing xsf files: 0.460515 s
Maximum CPU memory used: 3.940384 GB
Maximum GPU memory used: 0.000000 GB
======================================================================
3-2. Visualize the test results¶
energy_test_gpr = np.load("./energy_test_gpr.npy")
energy_test_reference = np.load("./energy_test_reference.npy")
force_test_gpr = np.load("./force_test_gpr.npy")
force_test_reference = np.load("./force_test_reference.npy")
unc_e_test_gpr = np.load("./unc_e_test_gpr.npy")
unc_f_test_gpr = np.load("./unc_f_test_gpr.npy")diff_per_atom = (energy_test_reference - energy_test_gpr) / 46
print(diff_per_atom.min(), diff_per_atom.max())-0.0027989477913133273 0.012551811322776737
fig = plt.figure(figsize=(7, 6))
ax2 = fig.subplots()
font_x = {'size': 20, 'color': 'black'}
font_y = {'size': 20, 'color': 'black'}
font_tick = {'size': 16, 'color': 'black'}
# The second plot
counts, bins = np.histogram(diff_per_atom, bins=30, range=(-0.005, 0.05))
ax2.hist(bins[:-1], bins, weights=counts, facecolor='dimgray', alpha=1.0, edgecolor='black')
ax2.set_xlabel("DFT $-$ GPR (eV/atom)", fontdict=font_x)
ax2.set_xlim([-0.03, 0.03])
x_labels = [round(label, 2) for label in ax2.get_xticks()]
ax2.set_xticks(x_labels)
ax2.set_xlim([-0.03, 0.03])
ax2.set_xticklabels(x_labels, fontdict=font_tick)
ax2.set_ylabel("Number of structures", fontdict=font_y)
ax2.set_ylim([0, 25])
y_labels = [int(label) for label in ax2.get_yticks()]
ax2.set_yticks(y_labels)
ax2.set_ylim([0, 25])
ax2.set_yticklabels(y_labels, fontdict=font_tick)
[x.set_linewidth(1.5) for x in ax2.spines.values()]
ax2.tick_params(bottom=False, top=True, left=True, right=True)
ax2.tick_params(labelbottom=True, labeltop=False, labelleft=True, labelright=False)
ax2.tick_params(direction='in', length=8, width=1.5)
ax2.grid(True, which='both', linestyle=':', linewidth=0.7, alpha=0.7)
fig.tight_layout()
plt.show()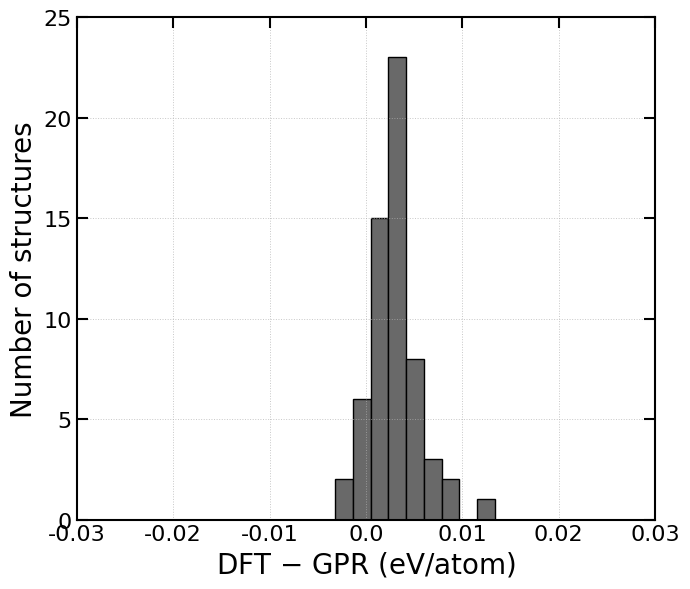
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import pearsonr
# --- Force error vs uncertainty ---
force_error = np.abs(force_test_gpr - force_test_reference).reshape(-1)
unc_f = unc_f_test_gpr.reshape(-1)
r_force, _ = pearsonr(force_error, unc_f)
plt.figure(figsize=(6, 5))
plt.scatter(unc_f, force_error, alpha=0.6, edgecolors='k')
plt.xlabel("Force uncertainty (eV/Å)")
plt.ylabel("Force absolute error (eV/Å)")
plt.title(f"Force Error vs Uncertainty\nPearson r = {r_force:.3f}")
plt.grid(True)
# plt.xlim(0.0, 1.0)
# plt.ylim(0, 2.0)
plt.tight_layout()
plt.show()