MatterGen in Practice: pretrained crystal diffusion
Source repo for Colab bootstrap and helper downloads: https://
MatterGen in Practice: pretrained crystal diffusion¶
This notebook is the first production-style continuation of crystal-diffusion-from-scratch.ipynb. Instead of building the whole model ourselves, we use a pretrained MatterGen checkpoint and focus on the scientific workflow around it: run generation, inspect the returned candidate pool, and decide what to screen next.
Aims¶
run a pretrained unconditional MatterGen model,
run a low-vs-high conditioned sweep with the same checkpoint,
parse the resulting structures into a common screening table,
inspect one explicit reverse-diffusion trajectory,
connect those diagnostics to realistic computational-chemistry decision making,
optionally borrow the next discovery step from the upstream tutorial by relaxing a shortlist of generated candidates.
Learning outcomes¶
By the end you should be able to:
explain what MatterGen is conditioning on in this notebook,
distinguish conditioning from post-generation screening,
read MatterGen outputs as a candidate pool rather than a single answer,
say when a pretrained crystal-diffusion workflow is more useful than a from-scratch teaching model.
Primary sources and upstream code¶
This notebook is organized around the official MatterGen resources and the Materials Project ecosystem it is designed to work with:
MatterGen repository: https://
github .com /microsoft /mattergen MatterGen paper: https://
www .nature .com /articles /s41586 -025 -08628-5 MatterGen commit pinned in this notebook:
a245cf2b7538eea6d873e6430b0e30c56d26c60eMaterials Project paper: Jain et al. (2013)
Materials Project API docs: https://
materialsproject .github .io /api/
The aim here is not to reproduce MatterGen pretraining. The aim is to learn how to run, inspect, and discuss a modern pretrained crystal-diffusion system in a classroom or research-notebook setting.
Table of Contents¶
Task for you
Before you run MatterGen, predict which quantities in this notebook are inputs to generation and which are only post hoc diagnostics.
Keep comparing to
crystal-diffusion-from-scratch.ipynb: what is now hidden inside the pretrained checkpoint, and what is still your responsibility as a scientist?Decide in advance what would count as a useful success signal: mode diversity, structural plausibility, a shifted distribution, or a clean trajectory.
1. Install prerequisites (Colab-friendly)¶
This section anchors the notebook to the notebooks/05-generative/ directory inside this tutorial repo, clones microsoft/mattergen locally, checks out the pinned commit a245cf2b7538, installs it into a repo-local Python 3.10 virtual environment, and sets up the analysis stack used later in the notebook.
The first run needs network access so the repo can be cloned and the checkpoints can be fetched when requested.
# @title
# Install MatterGen into a repo-local Python 3.10 environment and anchor all later paths to this notebook folder.
from pathlib import Path
import html
import os
import subprocess
import sys
DAY5_SOURCE_REPO_URL = "https://gitlab.com/cam-ml/tutorials.git"
DAY5_SOURCE_REPO_BRANCH = "main"
DAY5_COLAB_CLONE_CANDIDATES = [
Path("/content/tutorials"),
Path("/content/cam_ml_tutorials"),
Path("/content/camml-tutorials"),
]
def _running_in_colab():
try:
import google.colab # type: ignore
return True
except Exception:
return False
def _unique_paths(paths):
unique = []
seen = set()
for path in paths:
path = Path(path)
key = str(path)
if key not in seen:
seen.add(key)
unique.append(path)
return unique
def _iter_day5_search_roots():
cwd = Path.cwd().resolve()
roots = [cwd, *cwd.parents]
for clone_dir in DAY5_COLAB_CLONE_CANDIDATES:
roots.extend([clone_dir, clone_dir / "notebooks" / "05-generative"])
return _unique_paths(roots)
def _register_day5_notebook_root(notebook_root: Path):
notebook_root = notebook_root.resolve()
if str(notebook_root) not in sys.path:
sys.path.insert(0, str(notebook_root))
try:
os.chdir(notebook_root)
except OSError:
pass
return notebook_root
def ensure_day5_helpers_on_path():
for candidate in _iter_day5_search_roots():
for notebook_root in (candidate, candidate / "notebooks" / "05-generative"):
helper_dir = notebook_root / "gen_helpers"
if helper_dir.exists():
return _register_day5_notebook_root(notebook_root)
if _running_in_colab():
for clone_dir in DAY5_COLAB_CLONE_CANDIDATES:
notebook_root = clone_dir / "notebooks" / "05-generative"
if notebook_root.exists():
return _register_day5_notebook_root(notebook_root)
for clone_dir in DAY5_COLAB_CLONE_CANDIDATES:
if clone_dir.exists():
continue
clone_dir.parent.mkdir(parents=True, exist_ok=True)
print(
"Cloning the Day 5 tutorial repo from "
f"{DAY5_SOURCE_REPO_URL} into {clone_dir} so notebook helper modules are available..."
)
subprocess.run(
[
"git",
"clone",
"--depth",
"1",
"--branch",
DAY5_SOURCE_REPO_BRANCH,
DAY5_SOURCE_REPO_URL,
str(clone_dir),
],
check=True,
)
notebook_root = clone_dir / "notebooks" / "05-generative"
if notebook_root.exists():
return _register_day5_notebook_root(notebook_root)
raise FileNotFoundError(
"Could not find or clone notebooks/05-generative inside /content for this Colab session."
)
raise FileNotFoundError(
"Could not locate notebooks/05-generative/gen_helpers. If you are in Colab, rerun this cell so the repo can be cloned automatically."
)
GEN_HELPERS_ROOT = ensure_day5_helpers_on_path()
!pip install ase==3.25.0 ipywidgets
try:
import ipywidgets as widgets
except Exception:
widgets = None
try:
import google.colab # type: ignore
IN_COLAB = True
except Exception:
IN_COLAB = False
from IPython.display import display
from gen_helpers.discovery_workflow import (
build_relaxation_rows,
relax_atoms_with_mace,
show_before_after_relaxation,
)
from gen_helpers.mattergen_helpers import (
collect_mattergen_runs,
extract_mattergen_trajectory_preview,
inspect_generated_cif_archive,
plot_mattergen_diagnostics,
render_rank_table,
render_summary_table,
run_mattergen_generation as _run_mattergen_generation,
setup_mattergen_environment,
show_mattergen_gallery,
)
mattergen_env = setup_mattergen_environment()
NOTEBOOK_ROOT = mattergen_env["notebook_root"]
def run_mattergen_generation(*args, **kwargs):
return _run_mattergen_generation(mattergen_env, *args, **kwargs)
def format_widget_pre(text: str) -> str:
return f"<pre style='white-space:pre-wrap; margin:0'>{html.escape(text)}</pre>"
def bind_widget_state(controls, apply_fn):
state_holder = {"has_rendered": False, "last": None}
def refresh(change=None):
state = {name: control.value for name, control in controls.items()}
state_key = tuple((name, repr(value)) for name, value in state.items())
if state_holder["has_rendered"] and state_holder["last"] == state_key:
return
state_holder["has_rendered"] = True
state_holder["last"] = state_key
apply_fn(**state)
refresh()
for control in controls.values():
control.observe(refresh, names='value')
return refresh
# @title
import torch
import platform
print("Python:", platform.python_version())
print("PyTorch:", torch.__version__)
print("CUDA available:", torch.cuda.is_available())
if torch.cuda.is_available():
print("GPU:", torch.cuda.get_device_name(0))
2. Core ideas to keep in mind¶
Why generative modeling for computational chemistry?¶
Classical crystal discovery workflows are powerful but expensive. We want a model that can:
learn structural priors from known inorganic crystals,
propose novel candidate structures with plausible chemistry,
be steerable toward target properties or target crystal families.
Practical takeaway¶
The conditioning signal in this notebook is a scalar band-gap target plus a guidance factor.
Volume, atom count, formula diversity, and galleries are post-generation diagnostics. They help us ask whether the low- and high-target runs moved into different regions of crystal space, but they are not the target itself.
Useful scientific work begins after sampling: parse the pool, compare how the conditioned runs shifted, shortlist candidates, and only then move to relaxation or DFT.
Dataset note¶
The pretrained checkpoints in this notebook were not trained on molecular crystals, proteins, or arbitrary crystallographic databases. They come from the MatterGen training datasets built around Materials Project-style inorganic crystals, mainly the MP-20 / Alex-MP-20 family with at most 20 atoms per unit cell. That matters when you interpret failures or successes: the model prior is strongest in exactly that region of crystal space.
Official MatterGen Figures¶
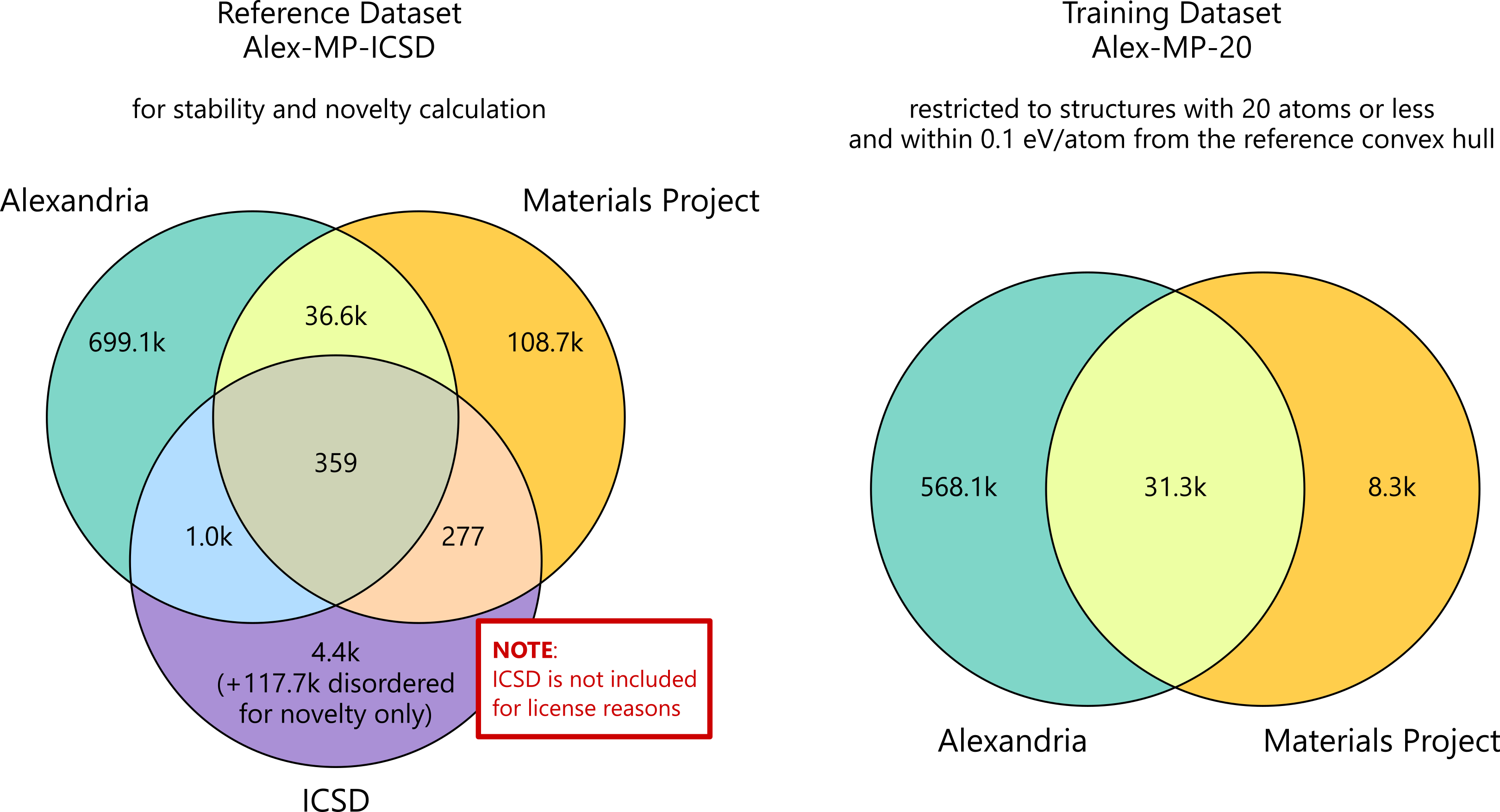
Source: official MatterGen repository asset assets/datasets_venn_diagram.png: https://
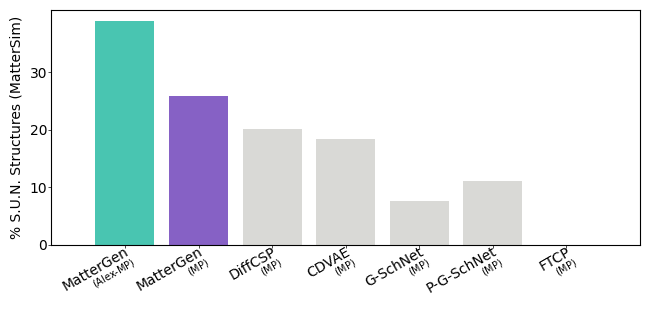
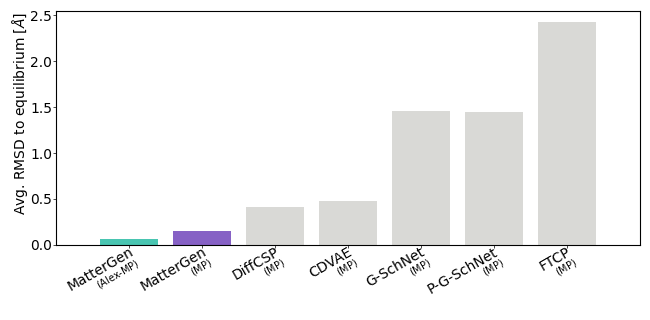
Source: official MatterGen repository benchmark figures benchmark/figures/metrics_sun.png and benchmark/figures/metrics_rmsd.png: https://
3. MatterGen architecture map (exploring the MatterGen code)¶
If you want to connect the notebook to the repository, start from the user-facing entrypoint and then work inward:
mattergen/scripts/generate.pyfor samplingmattergen/generator.pyfor the reverse-diffusion loopmattergen/property_embeddings.pyfor conditioningmattergen/diffusion/for losses and denoising blocksmattergen/scripts/finetune.pyfor conditioned adaptation
How to use this map¶
If you only want generated structures, learn the command-line entrypoint first.
If you want to change conditioning, inspect the property embedding path.
If you want to modify sampling behavior, start in
generator.py.
Mini exercise¶
Open two files side-by-side and trace one generated sample from condition to output file.
Suggested answer
mattergen/property_embeddings.py builds the conditioning signal, generator.py feeds that signal into the sampling loop, and scripts/generate.py turns the whole path into a runnable command-line workflow.
4. Unconditional generation with a pre-trained checkpoint¶
We start with the base model (mattergen_base) because it shows the simplest complete user flow: choose a checkpoint, run generation, inspect the files, and plot the samples.
What to notice¶
The command writes a results directory with
.extxyzsamples and recorded trajectories.--batch_sizeand--num_batchescontrol how much data you generate without changing the model.The output is not just one structure; it is a batch you can parse, compare, and visualize.
This is the direct crystal-diffusion workflow, wrapped as a reproducible model checkpoint.
Quick check: why try unconditional generation first?
Answer
It isolates the base crystal prior. Before we talk about guidance or property targets, we first want to see whether the pretrained model can generate plausible crystals at all.
MATTERGEN_FORCE_RERUN = globals().get('MATTERGEN_FORCE_RERUN', False) # @param {type:"boolean"}
MATTERGEN_UNCONDITIONAL_BATCH_SIZE = int(globals().get('MATTERGEN_UNCONDITIONAL_BATCH_SIZE', 3)) # @param {type:"integer"}
MATTERGEN_UNCONDITIONAL_NUM_BATCHES = int(globals().get('MATTERGEN_UNCONDITIONAL_NUM_BATCHES', 2)) # @param {type:"integer"}
MATTERGEN_RECORD_TRAJECTORIES = globals().get('MATTERGEN_RECORD_TRAJECTORIES', True) # @param {type:"boolean"}
def _mattergen_unconditional_summary() -> str:
lines = [
'MatterGen unconditional settings:',
f' batch_size: {MATTERGEN_UNCONDITIONAL_BATCH_SIZE}',
f' num_batches: {MATTERGEN_UNCONDITIONAL_NUM_BATCHES}',
f' record_trajectories: {MATTERGEN_RECORD_TRAJECTORIES}',
f' force_rerun: {MATTERGEN_FORCE_RERUN}',
]
if not IN_COLAB and widgets is not None:
lines.append('Adjust the widgets, then rerun this cell to launch a new unconditional batch.')
return '\n'.join(lines)
def _apply_mattergen_unconditional_controls(batch_size, num_batches, force_rerun, record_trajectories, announce: bool = True):
global MATTERGEN_UNCONDITIONAL_BATCH_SIZE, MATTERGEN_UNCONDITIONAL_NUM_BATCHES
global MATTERGEN_FORCE_RERUN, MATTERGEN_RECORD_TRAJECTORIES
MATTERGEN_UNCONDITIONAL_BATCH_SIZE = int(batch_size)
MATTERGEN_UNCONDITIONAL_NUM_BATCHES = int(num_batches)
MATTERGEN_FORCE_RERUN = bool(force_rerun)
MATTERGEN_RECORD_TRAJECTORIES = bool(record_trajectories)
summary = _mattergen_unconditional_summary()
if announce:
print(summary)
return summary
if IN_COLAB or widgets is None:
print(
_apply_mattergen_unconditional_controls(
MATTERGEN_UNCONDITIONAL_BATCH_SIZE,
MATTERGEN_UNCONDITIONAL_NUM_BATCHES,
MATTERGEN_FORCE_RERUN,
MATTERGEN_RECORD_TRAJECTORIES,
announce=False,
)
)
if not IN_COLAB and widgets is None:
print('Install `ipywidgets` to get Jupyter controls for this cell.')
else:
uncond_batch_widget = widgets.BoundedIntText(
value=MATTERGEN_UNCONDITIONAL_BATCH_SIZE,
min=1,
max=16,
description='Batch size:',
style={'description_width': '90px'},
layout=widgets.Layout(width='220px'),
)
uncond_num_batches_widget = widgets.BoundedIntText(
value=MATTERGEN_UNCONDITIONAL_NUM_BATCHES,
min=1,
max=12,
description='Batches:',
style={'description_width': '90px'},
layout=widgets.Layout(width='220px'),
)
uncond_record_widget = widgets.Checkbox(
value=bool(MATTERGEN_RECORD_TRAJECTORIES),
description='Record trajectories',
)
uncond_force_widget = widgets.Checkbox(
value=bool(MATTERGEN_FORCE_RERUN),
description='Force rerun',
)
uncond_help = widgets.HTML(
'<small>In Jupyter, update the controls below and rerun this cell when you want fresh MatterGen outputs.</small>'
)
uncond_status = widgets.HTML()
display(
widgets.VBox(
[
widgets.HBox([uncond_batch_widget, uncond_num_batches_widget]),
widgets.HBox([uncond_record_widget, uncond_force_widget]),
uncond_help,
uncond_status,
]
)
)
def _refresh_unconditional_controls(batch_size, num_batches, force_rerun, record_trajectories):
uncond_status.value = format_widget_pre(
_apply_mattergen_unconditional_controls(
batch_size,
num_batches,
force_rerun,
record_trajectories,
announce=False,
)
)
bind_widget_state(
{
'batch_size': uncond_batch_widget,
'num_batches': uncond_num_batches_widget,
'force_rerun': uncond_force_widget,
'record_trajectories': uncond_record_widget,
},
_refresh_unconditional_controls,
)
if not mattergen_env["mattergen_bin"].exists():
raise FileNotFoundError('MatterGen CLI not found. Run the install cell first.')
results_dir = run_mattergen_generation(
'results_unconditional',
pretrained_name='mattergen_base',
batch_size=MATTERGEN_UNCONDITIONAL_BATCH_SIZE,
num_batches=MATTERGEN_UNCONDITIONAL_NUM_BATCHES,
record_trajectories=MATTERGEN_RECORD_TRAJECTORIES,
force=MATTERGEN_FORCE_RERUN,
)
mattergen_run_info = {
'unconditional': {
'display_name': 'Unconditional base',
'pretrained_name': 'mattergen_base',
'target_label': 'none',
'condition_kind': 'unconditional',
'output_dir': results_dir,
}
}
print('\nDone. Files in output directory:')
for p in sorted(results_dir.iterdir()):
print(' ', p.name)
Inspect the raw files MatterGen wrote¶
Before moving on to conditioned generation, it is worth opening one of the files exactly as MatterGen exported it. Download the generated .zip in results_unconditional and inspect some of the .cif files. Do these crystal structures look sensible?
What to notice¶
_chemical_formula_sumgives the written composition._cell_length_*and_cell_angle_*encode the unit cell._space_group_name_H-M_altrecords the symmetry label stored in the CIF.the atom loop lists fractional coordinates and occupancies for each site.
from IPython.display import Markdown, display
cif_preview = inspect_generated_cif_archive(
mattergen_run_info["unconditional"]["output_dir"],
preview_index=0,
preview_lines=36,
)
print("MatterGen output folder:", mattergen_run_info["unconditional"]["output_dir"])
print("CIF archive:", cif_preview["archive_path"].name)
print("Archive members:", ", ".join(cif_preview["cif_names"]))
print("Showing raw text from:", cif_preview["preview_name"])
display(Markdown("```cif\n" + cif_preview["preview_excerpt"] + "\n```"))
print("Commentary:")
print("- MatterGen exports a proper crystallographic text file, not only arrays in Python memory.")
print("- The lattice constants and angles are explicit in the CIF header.")
print("- This sample is written as P1, which is useful to discuss: MatterGen does not explicitly enforce higher-level symmetries, but it can be up to downstream processing to identify and utilize any symmetries present in the generated structures.")
Task for you
In the CIF excerpt, point to the exact lines that tell you about composition, lattice geometry, and symmetry.
Compare
_chemical_formula_sumwith the atom list below it: do the stoichiometries agree?The sample is exported as
P1. Do you read that as “bad sample”, “low-confidence symmetry assignment”, or “needs further analysis”?
5. Property-conditioned generation: low and high band gap targets¶
A single conditioned run is useful, but a sweep is much more informative. Here we use the same fine-tuned MatterGen checkpoint twice, once with a low band-gap target and once with a high band-gap target.
Two things matter here:
the conditioned target is the band gap,
the downstream screening statistics we will compare later are ordinary crystal descriptors such as unit-cell volume, number of sites, and formula diversity.
Those screening views are not measuring target adherence directly. They are cheap structural diagnostics that help us see whether the conditional sweep is moving the generator into different regions of crystal space.
What to notice¶
We change the target value but keep the same conditioned checkpoint.
We keep the guidance factor fixed so the main thing changing is the requested target.
Looking at low and high targets side by side is the closest notebook-sized version of a conditioning study.
After generation, we treat the outputs as a candidate pool and ask whether the low and high targets changed the structural regime, not just whether one individual sample looks plausible.
Quick check: why is it useful to compare low and high targets from the same conditioned checkpoint?
Answer
It isolates the effect of the target value itself. If you change both the checkpoint and the target at the same time, it becomes harder to tell whether differences came from the learned conditional prior or from the requested condition.
# These are real MatterGen conditional runs using the same fine-tuned band-gap checkpoint
# at two very different target values.
MATTERGEN_LOW_BAND_GAP_TARGET = float(globals().get('MATTERGEN_LOW_BAND_GAP_TARGET', 0.2)) # @param {type:"number"}
MATTERGEN_HIGH_BAND_GAP_TARGET = float(globals().get('MATTERGEN_HIGH_BAND_GAP_TARGET', 4.0)) # @param {type:"number"}
MATTERGEN_CONDITIONAL_GUIDANCE = float(globals().get('MATTERGEN_CONDITIONAL_GUIDANCE', 2.0)) # @param {type:"number"}
MATTERGEN_CONDITIONAL_BATCH_SIZE = int(globals().get('MATTERGEN_CONDITIONAL_BATCH_SIZE', 2)) # @param {type:"integer"}
MATTERGEN_CONDITIONAL_NUM_BATCHES = int(globals().get('MATTERGEN_CONDITIONAL_NUM_BATCHES', 1)) # @param {type:"integer"}
MATTERGEN_CONDITIONAL_RECORD_TRAJECTORIES = globals().get('MATTERGEN_CONDITIONAL_RECORD_TRAJECTORIES', True) # @param {type:"boolean"}
MATTERGEN_CONDITIONAL_FORCE_RERUN = globals().get('MATTERGEN_CONDITIONAL_FORCE_RERUN', False) # @param {type:"boolean"}
def _mattergen_conditional_summary() -> str:
lines = [
'MatterGen conditional settings:',
f' low target (eV): {MATTERGEN_LOW_BAND_GAP_TARGET}',
f' high target (eV): {MATTERGEN_HIGH_BAND_GAP_TARGET}',
f' guidance: {MATTERGEN_CONDITIONAL_GUIDANCE}',
f' batch_size: {MATTERGEN_CONDITIONAL_BATCH_SIZE}',
f' num_batches: {MATTERGEN_CONDITIONAL_NUM_BATCHES}',
f' record_trajectories: {MATTERGEN_CONDITIONAL_RECORD_TRAJECTORIES}',
f' force_rerun: {MATTERGEN_CONDITIONAL_FORCE_RERUN}',
]
if not IN_COLAB and widgets is not None:
lines.append('Adjust the widgets, then rerun this cell to regenerate the conditioned sweeps.')
return '\n'.join(lines)
def _apply_mattergen_conditional_controls(low_gap, high_gap, guidance, batch_size, num_batches, record_trajectories, force_rerun, announce: bool = True):
global MATTERGEN_LOW_BAND_GAP_TARGET, MATTERGEN_HIGH_BAND_GAP_TARGET
global MATTERGEN_CONDITIONAL_GUIDANCE, MATTERGEN_CONDITIONAL_BATCH_SIZE, MATTERGEN_CONDITIONAL_NUM_BATCHES
global MATTERGEN_CONDITIONAL_RECORD_TRAJECTORIES, MATTERGEN_CONDITIONAL_FORCE_RERUN
MATTERGEN_LOW_BAND_GAP_TARGET = float(low_gap)
MATTERGEN_HIGH_BAND_GAP_TARGET = float(high_gap)
MATTERGEN_CONDITIONAL_GUIDANCE = float(guidance)
MATTERGEN_CONDITIONAL_BATCH_SIZE = int(batch_size)
MATTERGEN_CONDITIONAL_NUM_BATCHES = int(num_batches)
MATTERGEN_CONDITIONAL_RECORD_TRAJECTORIES = bool(record_trajectories)
MATTERGEN_CONDITIONAL_FORCE_RERUN = bool(force_rerun)
summary = _mattergen_conditional_summary()
if announce:
print(summary)
return summary
if IN_COLAB or widgets is None:
print(
_apply_mattergen_conditional_controls(
MATTERGEN_LOW_BAND_GAP_TARGET,
MATTERGEN_HIGH_BAND_GAP_TARGET,
MATTERGEN_CONDITIONAL_GUIDANCE,
MATTERGEN_CONDITIONAL_BATCH_SIZE,
MATTERGEN_CONDITIONAL_NUM_BATCHES,
MATTERGEN_CONDITIONAL_RECORD_TRAJECTORIES,
MATTERGEN_CONDITIONAL_FORCE_RERUN,
announce=False,
)
)
if not IN_COLAB and widgets is None:
print('Install `ipywidgets` to get Jupyter controls for this cell.')
else:
low_gap_widget = widgets.FloatSlider(value=MATTERGEN_LOW_BAND_GAP_TARGET, min=0.0, max=2.0, step=0.1, description='Low gap:', readout_format='.1f', continuous_update=False, style={'description_width': '90px'}, layout=widgets.Layout(width='280px'))
high_gap_widget = widgets.FloatSlider(value=MATTERGEN_HIGH_BAND_GAP_TARGET, min=2.0, max=6.0, step=0.1, description='High gap:', readout_format='.1f', continuous_update=False, style={'description_width': '90px'}, layout=widgets.Layout(width='280px'))
guidance_widget = widgets.FloatSlider(value=MATTERGEN_CONDITIONAL_GUIDANCE, min=0.5, max=5.0, step=0.25, description='Guidance:', readout_format='.2f', continuous_update=False, style={'description_width': '90px'}, layout=widgets.Layout(width='280px'))
conditional_batch_widget = widgets.BoundedIntText(value=MATTERGEN_CONDITIONAL_BATCH_SIZE, min=1, max=12, description='Batch size:', style={'description_width': '90px'}, layout=widgets.Layout(width='220px'))
conditional_num_batches_widget = widgets.BoundedIntText(value=MATTERGEN_CONDITIONAL_NUM_BATCHES, min=1, max=8, description='Batches:', style={'description_width': '90px'}, layout=widgets.Layout(width='220px'))
conditional_record_widget = widgets.Checkbox(value=bool(MATTERGEN_CONDITIONAL_RECORD_TRAJECTORIES), description='Record trajectories')
conditional_force_widget = widgets.Checkbox(value=bool(MATTERGEN_CONDITIONAL_FORCE_RERUN), description='Force rerun')
conditional_help = widgets.HTML('<small>In Jupyter, update the targets or guidance below and rerun this cell to launch a new conditioned sweep.</small>')
conditional_status = widgets.HTML()
display(widgets.VBox([
widgets.HBox([low_gap_widget, high_gap_widget]),
widgets.HBox([guidance_widget, conditional_batch_widget, conditional_num_batches_widget]),
widgets.HBox([conditional_record_widget, conditional_force_widget]),
conditional_help,
conditional_status,
]))
def _refresh_conditional_controls(low_gap, high_gap, guidance, batch_size, num_batches, record_trajectories, force_rerun):
conditional_status.value = format_widget_pre(
_apply_mattergen_conditional_controls(
low_gap,
high_gap,
guidance,
batch_size,
num_batches,
record_trajectories,
force_rerun,
announce=False,
)
)
bind_widget_state(
{
'low_gap': low_gap_widget,
'high_gap': high_gap_widget,
'guidance': guidance_widget,
'batch_size': conditional_batch_widget,
'num_batches': conditional_num_batches_widget,
'record_trajectories': conditional_record_widget,
'force_rerun': conditional_force_widget,
},
_refresh_conditional_controls,
)
MATTERGEN_CONDITIONAL_SPECS = {
'low_band_gap': {'display_name': 'Low band gap target', 'pretrained_name': 'dft_band_gap', 'properties': repr({'dft_band_gap': MATTERGEN_LOW_BAND_GAP_TARGET}), 'guidance': MATTERGEN_CONDITIONAL_GUIDANCE, 'batch_size': MATTERGEN_CONDITIONAL_BATCH_SIZE, 'num_batches': MATTERGEN_CONDITIONAL_NUM_BATCHES, 'target_label': f'dft_band_gap = {MATTERGEN_LOW_BAND_GAP_TARGET:.2f} eV', 'condition_kind': 'scalar target'},
'high_band_gap': {'display_name': 'High band gap target', 'pretrained_name': 'dft_band_gap', 'properties': repr({'dft_band_gap': MATTERGEN_HIGH_BAND_GAP_TARGET}), 'guidance': MATTERGEN_CONDITIONAL_GUIDANCE, 'batch_size': MATTERGEN_CONDITIONAL_BATCH_SIZE, 'num_batches': MATTERGEN_CONDITIONAL_NUM_BATCHES, 'target_label': f'dft_band_gap = {MATTERGEN_HIGH_BAND_GAP_TARGET:.2f} eV', 'condition_kind': 'scalar target'},
}
for label, spec in MATTERGEN_CONDITIONAL_SPECS.items():
out_dir = run_mattergen_generation(f'results_{label}', pretrained_name=spec['pretrained_name'], batch_size=spec['batch_size'], num_batches=spec['num_batches'], record_trajectories=MATTERGEN_CONDITIONAL_RECORD_TRAJECTORIES, properties=spec['properties'], guidance=spec['guidance'], force=MATTERGEN_CONDITIONAL_FORCE_RERUN)
mattergen_run_info[label] = {**spec, 'output_dir': out_dir}
print('\nMatterGen run registry:')
for label, spec in mattergen_run_info.items():
print(f"- {label}: {spec['display_name']} -> {spec['output_dir']}")
6. Parse MatterGen outputs into Python objects¶
Now we gather the unconditional run and the low/high conditioned sweeps into one Python-side structure so we can compare them consistently.
# Keep the notebook analysis stack aligned with the real MatterGen environment.
!python -m pip install --force-reinstall numpy==1.26.4 scipy==1.13.1 ase==3.25.0 matplotlib==3.8.4 pymatgen
mattergen_atoms, mattergen_rows, summary_rows = collect_mattergen_runs(mattergen_run_info)
print("\nMatterGen run summary:")
render_summary_table(summary_rows)
print("Reminder: the conditioned target here is band gap, while volume, atom count, and formula diversity are downstream structural diagnostics.")
Inspect a few unconditional generations by themselves¶
Before mixing the unconditional and conditioned runs together, pause and look at several unconditional samples on their own. This gives you a better feel for the base crystal prior before you ask whether the band-gap targets changed anything.
unconditional_rows = [row for row in mattergen_rows if row["label"] == "unconditional"]
render_rank_table(
"First unconditional MatterGen candidates to inspect closely",
unconditional_rows[:4],
["formula", "n_sites", "volume"],
)
show_mattergen_gallery(
mattergen_atoms,
unconditional_rows[:4],
"First unconditional MatterGen generations",
columns=2,
)
Task for you
After reading the raw CIF and the unconditional gallery, choose one sample you would screen first and one sample you would reject first. Explain why.
Before you inspect the conditioning-shift plots, predict which descriptor should move most strongly when you switch from the low to the high band-gap target: atom count, volume, or formula diversity.
Ask yourself which of these diagnostics would still be useful if the target were something other than band gap.
7. Conditioning shifts, galleries, and lightweight screening¶
This section checks whether the low and high band-gap targets produce visibly different structural distributions.
What to notice¶
We are conditioning on band gap but screening on structure.
Volume, atom count, and formula diversity are not substitutes for proper stability analysis, but they are cheap and immediately informative.
The most useful question is not “which sample looks most extreme?” but “did the low and high targets shift the generated pool in a consistent direction?”
The shortlist tables are intentionally practical: this is close to the first triage step you might do before relaxation or DFT.
palette = {
"unconditional": "#4c78a8",
"low_band_gap": "#54a24b",
"high_band_gap": "#e45756",
}
analysis = plot_mattergen_diagnostics(mattergen_run_info, mattergen_rows, palette=palette)
volume_sorted_rows = analysis["sorted_by_volume"]
representative_rows = analysis["representative_rows"]
shift_rows = analysis["shift_rows"]
render_rank_table(
"How the low-vs-high sweep shifts simple structural diagnostics",
shift_rows,
["run", "target", "mean_n_sites", "delta_mean_n_sites", "mean_volume", "delta_mean_volume", "unique_formulas"],
)
render_rank_table(
"Smallest-volume MatterGen candidates across all runs",
volume_sorted_rows[:4],
["display_name", "target_label", "formula", "n_sites", "volume"],
)
render_rank_table(
"Largest-volume MatterGen candidates across all runs",
list(reversed(volume_sorted_rows[-4:])),
["display_name", "target_label", "formula", "n_sites", "volume"],
)
show_mattergen_gallery(
mattergen_atoms,
representative_rows,
"Representative MatterGen samples across the unconditional and band-gap-conditioned runs",
columns=3,
)
show_mattergen_gallery(
mattergen_atoms,
volume_sorted_rows[:3],
"Smallest-volume MatterGen candidates across all runs",
columns=3,
)
show_mattergen_gallery(
mattergen_atoms,
list(reversed(volume_sorted_rows[-3:])),
"Largest-volume MatterGen candidates across all runs",
columns=3,
)
Optional extension: shortlist relaxation with MACE¶
We can go one step beyond generation and treat the outputs as candidates to relax and validate. Here we relax one representative structure from each MatterGen run and compare before/after geometries.
!python -m pip install mace-torch==0.3.14 torch-sim-atomistic==0.3.0 pymatgenMATTERGEN_RELAX_RUN = True
MATTERGEN_RELAX_DEVICE = "cuda"
MATTERGEN_RELAX_MODEL_SIZE = "small" # small, medium
mattergen_relax_rows = []
seen_labels = set()
for row in representative_rows:
if row['label'] not in seen_labels:
seen_labels.add(row['label'])
mattergen_relax_rows.append(row)
mattergen_relax_atoms = [mattergen_atoms[row['label']][row['local_index']] for row in mattergen_relax_rows]
mattergen_relax_labels = [f"{row['display_name']} | {row['formula']}" for row in mattergen_relax_rows]
if not MATTERGEN_RELAX_RUN:
print('Set MATTERGEN_RELAX_RUN=True and rerun this cell to relax one representative candidate from each MatterGen run.')
else:
try:
mattergen_relaxation = relax_atoms_with_mace(
mattergen_relax_atoms,
device=MATTERGEN_RELAX_DEVICE,
model_size=MATTERGEN_RELAX_MODEL_SIZE,
)
except ImportError as exc:
print('Optional relaxation dependencies are missing in this Python environment.')
print('Set MATTERGEN_OPTIONAL_VALIDATION_INSTALL=True and rerun the install cell first.')
print(f'Details: {exc}')
else:
mattergen_relaxation_rows = build_relaxation_rows(
mattergen_relax_atoms,
mattergen_relaxation['relaxed_atoms'],
mattergen_relaxation['energies'],
labels=mattergen_relax_labels,
)
render_rank_table(
'MatterGen shortlist after MACE relaxation',
mattergen_relaxation_rows,
['label', 'formula_before', 'formula_after', 'n_sites', 'energy_eV', 'volume_before', 'volume_after', 'delta_volume'],
)
show_before_after_relaxation(
mattergen_relax_atoms,
mattergen_relaxation['relaxed_atoms'],
mattergen_relax_labels,
'MatterGen shortlist before and after relaxation',
save_path=NOTEBOOK_ROOT / 'mattergen_relaxation_shortlist.png',
)
8. 3D visualization of a generated crystal¶
Before looking at a full diffusion path, it helps to inspect one finished sample by itself.
If you printed gen_0.cif above, compare the raw crystallographic text with the rendered structure here. This is the same kind of translation you would do in a real workflow when moving between file outputs and visual inspection.
What to notice¶
A single final sample lets you ask the simplest question first: does the decoded crystal look plausible?
Looking at two viewing angles helps separate lattice shape from atomic arrangement.
This is a structure-level check; the trajectory section below is a process-level check.
import matplotlib.pyplot as plt
from ase.io import read
from ase.visualize.plot import plot_atoms
from IPython.display import Image, display
preview_dir = (NOTEBOOK_ROOT / 'mattergen_figures').resolve()
preview_dir.mkdir(parents=True, exist_ok=True)
unconditional_extxyz = mattergen_run_info['unconditional']['output_dir'] / 'generated_crystals.extxyz'
if not unconditional_extxyz.exists():
raise FileNotFoundError(f'Could not find {unconditional_extxyz}. Run the unconditional MatterGen cell first.')
unconditional_atoms = read(unconditional_extxyz, index=':')
selected_sample_index = int(globals().get('selected_sample_index', 0)) # @param {type:"integer"}
selected_sample_index = max(0, min(selected_sample_index, len(unconditional_atoms) - 1))
def _render_mattergen_sample(sample_index: int, info_widget=None, image_widget=None):
global selected_sample_index
selected_sample_index = int(sample_index)
selected_atoms = unconditional_atoms[selected_sample_index]
preview_path = preview_dir / f'mattergen_unconditional_sample_{selected_sample_index:03d}.png'
summary = '\n'.join([
f'Showing unconditional sample {selected_sample_index} from {unconditional_extxyz}',
f'Formula: {selected_atoms.get_chemical_formula()}',
f'Number of atoms: {len(selected_atoms)}',
f'Volume: {selected_atoms.get_volume():.2f} ų',
])
fig, axes = plt.subplots(1, 2, figsize=(10, 4), squeeze=False, facecolor='white')
for ax, rotation, title in zip(axes[0], ['20x,30y,0z', '90x,0y,0z'], ['Perspective view', 'Side view']):
plot_atoms(selected_atoms, ax, rotation=rotation, radii=0.35, show_unit_cell=2)
ax.set_title(title, fontsize=10)
ax.set_axis_off()
plt.tight_layout()
fig.savefig(preview_path, dpi=180, bbox_inches='tight', facecolor='white')
plt.close(fig)
if info_widget is None:
print(summary)
else:
info_widget.value = format_widget_pre(summary)
if image_widget is None:
display(Image(filename=str(preview_path)))
else:
image_widget.value = preview_path.read_bytes()
if IN_COLAB or widgets is None:
_render_mattergen_sample(selected_sample_index)
if not IN_COLAB and widgets is None:
print('Install `ipywidgets` to browse different generated samples with a slider in Jupyter.')
else:
sample_index_widget = widgets.IntSlider(value=selected_sample_index, min=0, max=len(unconditional_atoms) - 1, step=1, description='Sample:', continuous_update=False, style={'description_width': '70px'}, layout=widgets.Layout(width='420px'))
sample_help = widgets.HTML('<small>Move the slider to inspect a different unconditional MatterGen sample.</small>')
sample_info = widgets.HTML()
sample_image = widgets.Image(format='png', layout=widgets.Layout(width='100%'))
display(widgets.VBox([sample_index_widget, sample_help, sample_info, sample_image]))
bind_widget_state(
{'sample_index': sample_index_widget},
lambda sample_index: _render_mattergen_sample(sample_index, info_widget=sample_info, image_widget=sample_image),
)
Quick check: why is it useful to inspect a final sample separately from a diffusion trajectory?
Answer
They answer different questions. A final-sample plot asks whether the decoded crystal itself looks plausible, while a trajectory plot asks how the model moved from noise toward that sample and whether the denoising path looks smooth or erratic.
MatterGen trajectory archive walkthrough¶
MatterGen can also record the reverse-diffusion path, not just the final crystals. In this section we unpack that archive, select one trajectory file, and follow that single candidate from noisy start to final structure.
Unpack the recorded trajectory archive and choose one path¶
from IPython.display import Image, display
trajectory_preview = extract_mattergen_trajectory_preview(mattergen_run_info, NOTEBOOK_ROOT)
display(Image(filename=str(trajectory_preview["preview_path"])))
display(Image(filename=str(trajectory_preview["gif_path"])))
What to notice¶
Every panel above comes from the same extracted trajectory file.
Early frames should look much noisier or less chemically stable than the late frames.
The exact intermediate chemistry can fluctuate during denoising; that is fine. The main question is whether one candidate path becomes progressively more structured.
9. Exercises¶
Low vs high targets: Why is a low-vs-high target sweep more informative than a single conditioned run?
Suggested answer
A sweep shows directionality. You are no longer asking only whether conditioning changes the samples, but whether changing the target moves the outputs in a consistent way.
Guidance sweep: Repeat the band-gap runs with guidance factors
[0.0, 1.0, 2.0, 4.0]and compare diversity versus how far the structural statistics move.
Suggested answer
Higher guidance should usually make the conditional signal stronger, but the samples may become less diverse or less realistic if the guidance is too aggressive.
Screening logic: Why is it sensible to compare atom count, cell volume, and formula diversity before running a more expensive relaxation or DFT workflow?
Suggested answer
Those statistics are cheap to compute and can quickly tell you whether the conditioned runs really shifted the generated pool before you spend time on more expensive downstream analysis.
Property vs structure: Which part of this MatterGen demo reflects the conditioning signal, and which part reflects the screening workflow?
Suggested answer
The band-gap target and guidance are the conditioning signal. The volume, atom-count, formula-diversity, gallery, and shortlist steps are the screening workflow applied after generation.
Raw CIF reading: Which lines in the exported CIF would you read first to sanity-check a MatterGen output, and why is seeing
P1not automatically a failure?
Suggested answer
Start with the written formula, the cell lengths and angles, and the stored space-group label, then check the atom-site loop to confirm the stoichiometry and coordinates are sensible. P1 is not automatically a failure because it may reflect a low-symmetry candidate, a noisy unrelaxed structure, or a conservative symmetry assignment before any downstream cleanup or relaxation.
Optional validation funnel: What does the MACE-relaxation step add that the raw generation plots cannot tell you by themselves?
Suggested answer
Relaxation tests whether a generated candidate settles into a more physically reasonable local minimum. The raw plots show distribution shifts, but they do not tell you how the geometry responds when you let the structure move downhill in energy.
10. Troubleshooting (Colab)¶
Out of memory: reduce
--batch_size.Slow runtime: ensure GPU runtime is selected. MatterGen’s reverse process is long enough that conditioned sweeps can feel slow on CPU.
Package conflicts: restart runtime and rerun setup cells.
MatterGen install problems: confirm network access, the
mattergen_repoclone, and rerun the repo-local venv install cell.Checkpoint download delays: the first call to a new pretrained checkpoint may take time while files are fetched from Hugging Face.
Analysis cells look stale: rerun the MatterGen parsing and visualization cells after generating fresh outputs.
You want a fresh run: set
MATTERGEN_FORCE_RERUN = Truein the generation cell before rerunning the notebook.
Summary¶
MatterGen gives us four complementary views of crystal diffusion in practice:
an unconditional crystal prior,
a low-vs-high conditioned sweep from the same checkpoint,
an optional shortlist-relaxation step borrowed from the upstream discovery tutorial,
and one explicit reverse-diffusion path for a single candidate crystal.
If you can explain which parts of this notebook are conditioning and which parts are screening, you already understand the most important scientific distinction in the workflow.
The next notebook, chemeleon-crystals.ipynb, keeps the same broad diffusion family but reorganizes the user-facing workflow around task choice: open-ended DNG or formula-conditioned CSP.
- Jain, A., Ong, S. P., Hautier, G., Chen, W., Richards, W. D., Dacek, S., Cholia, S., Gunter, D., Skinner, D., Ceder, G., & Persson, K. A. (2013). Commentary: The Materials Project: A materials genome approach to accelerating materials innovation. APL Materials, 1(1). 10.1063/1.4812323